 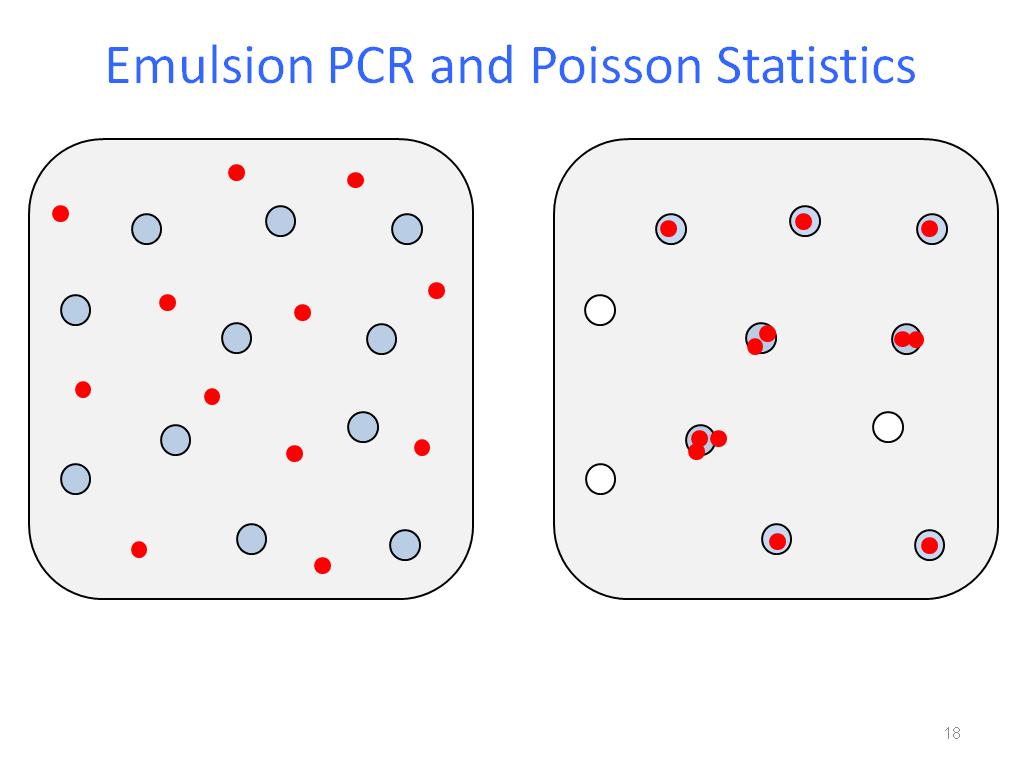
Instead, the agarose droplets themselves function as beads, Yang explains. 
Solution: Yang’s group turned to agarose-based PCR, but jettisoned the microbeads. “Running micrometer-sized beads in micrometer channels is prone to clogging,” he says, and, because beads have to squeeze through the channels, “the run rate is really slow.” He hoped to avoid the drawbacks of using microbeads to isolate molecules into discrete reactions. Yang wanted to streamline the aptamer screening process so that only quality aptamers were sequenced. The last step, individually analyzing each aptamer’s binding, kills any flickering hope of high-throughput aptamer discovery. These candidate aptamers are sequenced-an expensive step-and their sequences compared to identify aptamers that might bind well. It involves cloning candidate aptamer sequences into plasmids, transfecting these into bacteria, growing up the bacteria, and picking clones for aptamer analysis. After amplifying a library of possible sequences, actually generating the aptamers is quite time-consuming. Problem: Designing aptamers is generally a laborious, inefficient process. Aptamers that bind Shp2 and block its activity may have potential for treating Shp2-dependent cancers. Project: Efficiently select aptamers-segments of DNA with specific binding properties-against targets such as the cancer biomarker called SH2 domain–containing protein tyrosine phosphatase 2 (Shp2). Laboratory: Chaoyong James Yang, Lu Jiaxi Professor, Department of Chemical Biology, Xiamen University, Fujian, China Here are three case studies of researchers stretching ePCR’s capabilities. Tiemann-Boege and Novak are two of many scientists tweaking ePCR to maximize its benefits while minimizing its weaknesses. With standard PCR, you can amplify 1 to 2 kilobases,” Tiemann-Boege explains, but “the maximum in ePCR is 150 base pairs.” Commercial ePCR can also be a problem for anyone needing reaction volumes larger than a picoliter, prompting scientists to continue searching for creative solutions, says Richard Novak, of the University of California, Berkeley. In ePCR, “the product length is very short. Unfortunately, says Irene Tiemann-Boege, who studies genome recombination at Johannes Kepler University in Linz, Austria, emulsion PCR (ePCR) has a serious drawback for those hoping to cast a careful eye over the entire genome. (See “Prime Time for Digital PCR,” The Scientist, December 2011.) A plus is that the large number of individual reactions minimizes cross-contamination between products, but the greatest advantage is the ability to turn a tedious experiment into a high-throughput extravaganza. The amplification reaction can be carried out in many of the digital PCR machines on the market. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
